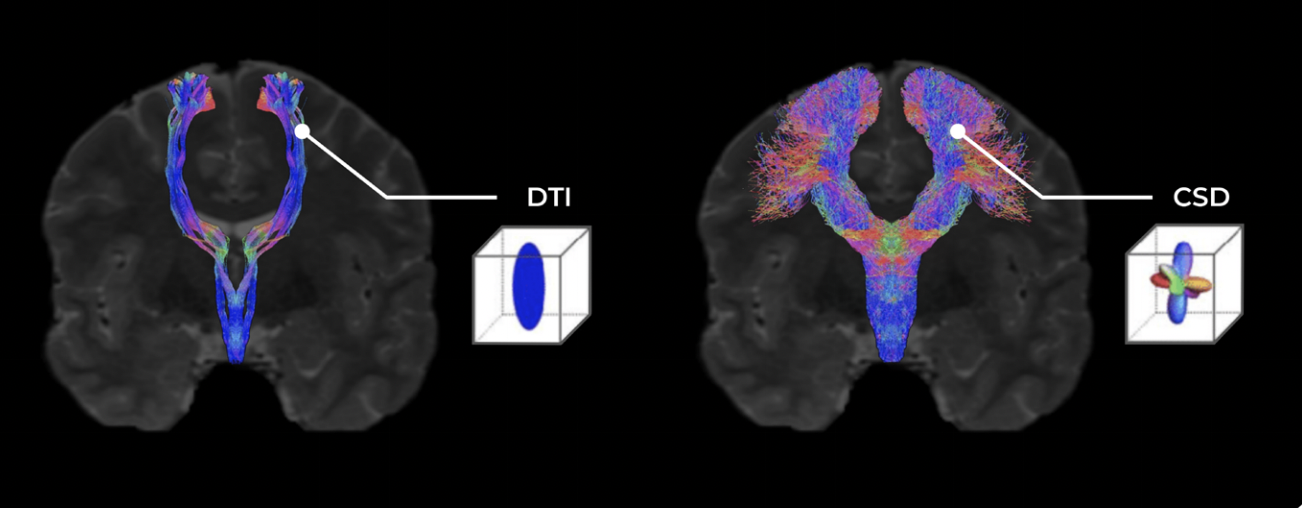
Imagine white matter tracts as the highways which connect the brain's major cities, or in this case, functional gray matter regions together. Previous and conventional methods of resolving DWI, such as DTI, considered these highways as spatially independent, connecting two cities together as a single road. However, highways typically also involve off-ramps, overpasses, and adjacent streets in close proximity. White matter structures within the cortex too hold greater complexity that cannot be accurately represented by a single directional value.
The primary limitation of previous techniques is that only one value, or tensor, representing the average direction of white matter tracts within a cubic voxel was able to be generated3,6. Whilst this assumption may enable the visualization of larger structures such as the Arcuate Fasciculus of the language system and Spinothalamic Tract of the sensorimotor system, it neglects that many smaller white matter structures originate as branches from or cross paths with these major structures, and would therefore be averaged out by the overarching diffusivity direction within voxels where an overlap is present6.
Arguably the greatest challenge in tractography analyses to date is resolving these high-density regions of white matter which include a number of crossing or touching fibers. Whilst a number of techniques have been developed to address this issue, the most superior, yet computationally demanding solution is “Constrained Spherical Deconvolution” (CSD).
Instead of an ‘average water diffusivity' value, CSD tractography generates a ‘fiber orientation distribution’ value for each voxel of neural tissue collected in the DWI series. This distribution value instead provides information on: a) the dominant diffusion signal within a voxel (major tracts), b) other independent direction signals also present within a voxel (fibers that cross under or above the dominant tract, and c) diffusion signals which branch from the dominant signal (fibers which branch or make contact with the major tract)7-9.

.png?width=1600&height=1080&name=Image_%20Hero-Corticospinal-Parcels%20(A).png)
%20(1).png?width=748&height=421&name=CSD%20(1)%20(1).png)